CopciAB_373418
Coprinopsis cinerea A43mutB43mut pab1-1 #326
General data
| Systematic name | CopciAB_373418 | Strain | Coprinopsis cinerea Okayama 7 |
|---|---|---|---|
| Standard name | - | Synonyms | 373418 |
| Uniprot id | Functional description | Glycoside hydrolase family 3 protein | |
| Location | scaffold_2:2957523..2960545 | Strand | + |
| Gene length (nt) | 3023 | Transcript length (nt) | 2320 |
| CDS length (nt) | 2211 | Protein length (aa) | 736 |
Reciprocal best hits in model fungi
| Strain name | Gene / Protein name |
|---|---|
Orthologs in mushroom models
| Strain name | Gene / Protein name | Pident | E-value | Bits |
|---|---|---|---|---|
| Hypsizygus marmoreus strain 51987-8 | Hypma_RDB26964 | 74.4 | 0 | 1118 |
| Agrocybe aegerita | Agrae_CAA7265302 | 71.4 | 0 | 1100 |
| Agaricus bisporus var bisporus (H97) | Agabi_varbisH97_2_217305 | 68 | 0 | 1051 |
| Agaricus bisporus var. burnettii JB137-S8 | Agabi_varbur_1_69320 | 68 | 0 | 1051 |
| Flammulina velutipes | Flave_chr11AA01097 | 64 | 2.658E-304 | 985 |
| Pleurotus eryngii ATCC 90797 | Pleery1_1398709 | 66 | 1.39E-304 | 957 |
| Pleurotus ostreatus PC9 | PleosPC9_1_61232 | 65.5 | 6.744E-304 | 949 |
| Pleurotus ostreatus PC15 | PleosPC15_2_1034680 | 65.2 | 8.169E-304 | 943 |
| Schizophyllum commune H4-8 | Schco3_2538080 | 61 | 3.668E-300 | 927 |
| Lentinula edodes W1-26 v1.0 | Lentinedodes1_20658 | 61.3 | 9.164E-292 | 917 |
| Lentinula edodes NBRC 111202 | Lenedo1_1166622 | 61.1 | 1.005E-295 | 914 |
| Auricularia subglabra | Aurde3_1_1406150 | 51.7 | 3.599E-237 | 745 |
Expression
| Name | Summary | Attribution | Assay type | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Carbon metabolism | Transcriptional changes along alternative morphogenetic paths of C. cinerea | Xie et al. 2020 | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Complex carbon sources 1 (long incubation) | Expression profiling on diverse complex carbon sources. | Hegedus et al. | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Coprinopsis challenged with bacteria | Transcriptional response to Bacillus subtilis and Escherichia coli. | Kombrink et al. 2018 | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Early development 1 | Time series expression profiling of basidiospore and oidium germination | Hegedus et al. | QuantSeq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Early development 2 | Time series expression profiling of mycelial growth, light induction and fruiting body initiation. | Hegedus et al. | QuantSeq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Fruiting body development | RNA-Seq analysis of fruiting body development | Krizsan et al. 2019 | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Nitrogen sources (12h incubation) | Expression profiling on diverse nitrogen sources. | Hegedus et al. | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Plant biomass (12h incubation) | Expression profiling on diverse plant biomasses as carbon sources. | Hegedus et al. | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Simple and complex carbon and nitrogen sources | Expression profiling on diverse complex carbon and nitrogen sources. | Hegedus et al. | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Simple and complex carbon sources (12h incubation) | Expression profiling on diverse complex carbon sources. | Hegedus et al. | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Starvation induced by water agar transfer | Expression profiling on starvation induced by transfer to water agar | Hegedus et al. | QuantSeq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Straw time series | Time series expression profiling on straw as a carbon source | Hegedus et al. | RNA-Seq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stress conditions 1 | Expression profiling under various stress conditions | Hegedus et al. | QuantSeq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stress conditions 2 | Expression profiling under various stress conditions | Hegedus et al. | QuantSeq | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
General data
| Systematic name | - |
|---|---|
| Protein id | CopciAB_373418.T0 |
| Description | Glycoside hydrolase family 3 protein |
Annotation summary
Conserved domains
| Analysis | Signature accession | Signature description | InterPro Accession | Start | End |
|---|---|---|---|---|---|
| Pfam | PF00933 | Glycosyl hydrolase family 3 N terminal domain | IPR001764 | 46 | 309 |
| Pfam | PF01915 | Glycosyl hydrolase family 3 C-terminal domain | IPR002772 | 362 | 623 |
| Pfam | PF14310 | Fibronectin type III-like domain | IPR026891 | 658 | 725 |
SignalP
| Prediction | Start | End | Score |
|---|---|---|---|
| SP(Sec/SPI) | 1 | 19 | 0.994 |
Transmembrane domains
| Domain n | Start | End | Length |
|---|---|---|---|
| No records | |||
InterPro
| Accession | Description |
|---|---|
| IPR017853 | Glycoside hydrolase superfamily |
| IPR026891 | Fibronectin type III-like domain |
| IPR001764 | Glycoside hydrolase, family 3, N-terminal |
| IPR036881 | Glycoside hydrolase family 3 C-terminal domain superfamily |
| IPR036962 | Glycoside hydrolase, family 3, N-terminal domain superfamily |
| IPR002772 | Glycoside hydrolase family 3 C-terminal domain |
| IPR013783 | Immunoglobulin-like fold |
GO
| Go id | Term | Ontology |
|---|---|---|
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds | MF |
| GO:0005975 | carbohydrate metabolic process | BP |
KEGG
| KEGG Orthology |
|---|
| K05349 |
EggNOG
| COG category | Description |
|---|---|
| G | Glycoside hydrolase family 3 protein |
CAZy
| Class | Family | Subfamily |
|---|---|---|
| GH | GH3 |
Transcription factor
| Group |
|---|
| No records |
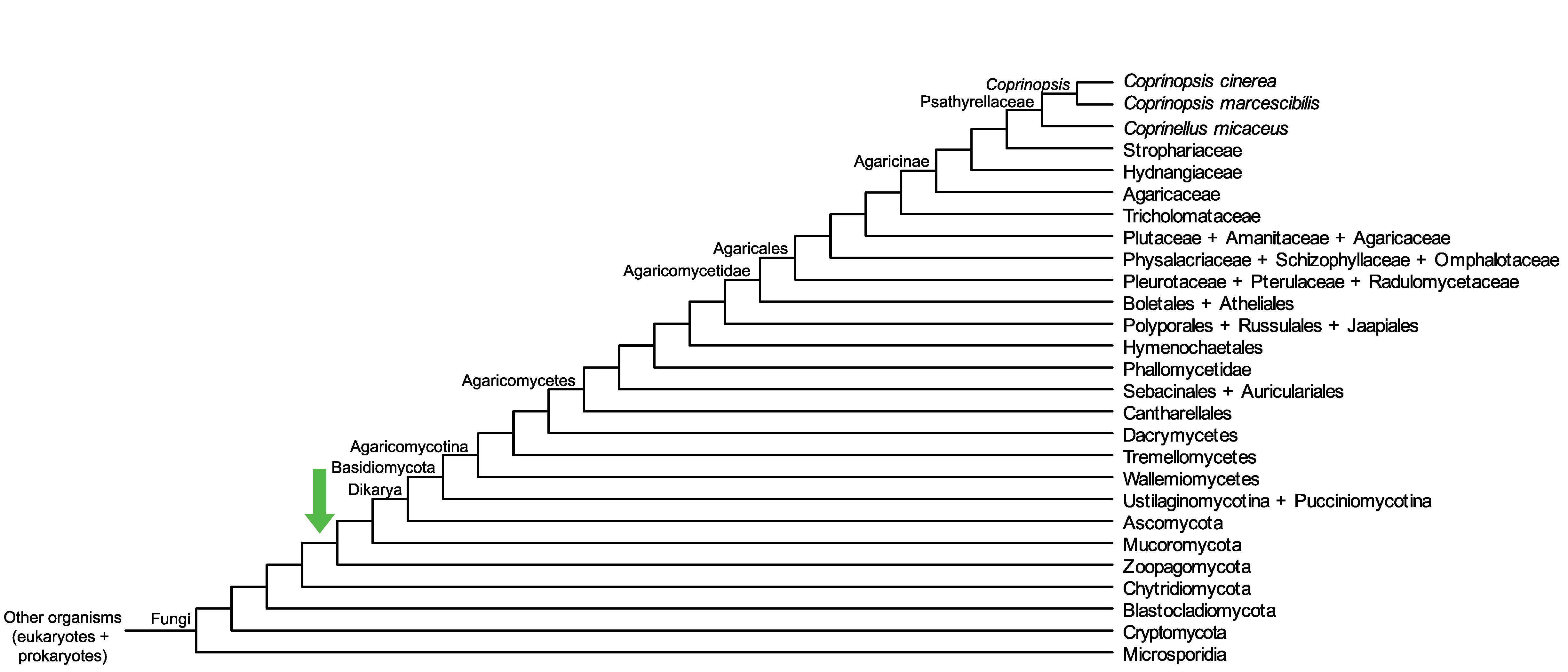
Conservation of CopciAB_373418 across fungi.
Arrow shows the origin of gene family containing CopciAB_373418.
Protein
| Sequence id | CopciAB_373418.T0 |
|---|---|
| Sequence |
>CopciAB_373418.T0 MRSSFSLLVLLAILQGTVARRTWDEAYDLANNVVSQMTLDEKLGIVRGTGQLNPNRRCVGDTKAVPRLGIPSICF ADGPAGMRLVKETTGFPTGINAAATFSRRLMRERGEALGREFRGKGSHVFLGPSVDVMRNPKGGRAWEAFGPDPY LNGEGAYETIMGVQSVGVQACIKHLIGNTQEHWRYGYSANMDDRTLHEMYFYPFLRSIEADVSSVMCAYNRFNGT SSCQNAGLLGDNGILRKAGFKGYVVSDWGATHDSAEVNANNGLDMEQPGDYIVVGGGVFGGLKDAVNKGRVSQRR LNEMVARILAPWYYLEQDSPDYPETNFDVQRRDGSGPKNLRVNVRSEEHTALAREIASASSVLLKNARSTTDGTP SGITTRGLPLSKDRIKSIAVIGQDARQLNQNCSEMGECNDGTMVVGWGSGSINFDHVVPPIDAIKSFTSDTATIT ESLTNNLDDGVRAARGKDAALVFVNAASGELAFYTVVHGNMGDRNDLDLWWRGGSLIERVAAVNNNTIVVVHSVG PVYMPWSTHPNITGIIYAGAPGEQTGPSIVDILYGNYNPSGKLPFSIADNEAAYGTTIVYNSLGFPDIDFTEKLL LDYRYMDEKNIRPRFEFGFGLSYTTFGYSNLAITRDGPGFQVSFTVANTGAFAGAEKPQLYLAYPPNAGQPKKVL RGFEEAILEVGASTRVTINLSERDMSVWDVPSQSYVRLPGVYTIYVGASSLDTRLQGTVTI |
| Length | 736 |
Coding
| Sequence id | CopciAB_373418.T0 |
|---|---|
| Sequence |
>CopciAB_373418.T0 ATGCGTTCTTCCTTCTCATTACTTGTGTTGCTCGCCATTCTCCAGGGCACTGTAGCCAGGAGAACATGGGACGAA GCTTATGACCTAGCCAACAATGTGGTGAGCCAGATGACCTTGGACGAGAAGTTGGGAATCGTCAGGGGCACAGGC CAACTCAACCCTAACCGTCGTTGTGTTGGTGATACCAAGGCTGTTCCCCGACTGGGCATCCCGTCAATCTGTTTC GCAGATGGCCCCGCAGGAATGCGTCTCGTAAAAGAGACGACCGGCTTCCCTACTGGCATTAACGCCGCAGCAACC TTCAGTCGTCGCCTCATGAGAGAGCGTGGTGAGGCTTTGGGTCGGGAGTTCCGAGGCAAAGGATCACACGTATTC TTGGGCCCATCCGTGGACGTTATGAGGAACCCCAAGGGAGGCCGTGCTTGGGAAGCCTTTGGTCCTGATCCTTAT CTGAACGGAGAAGGCGCTTATGAGACTATCATGGGCGTCCAGAGCGTTGGAGTTCAAGCCTGCATCAAGCACCTC ATCGGCAATACCCAGGAGCACTGGAGATATGGTTACAGCGCCAACATGGACGACAGAACACTTCACGAGATGTAC TTTTATCCATTCCTTCGAAGTATTGAGGCCGACGTCTCGTCCGTCATGTGCGCTTACAATCGCTTCAACGGCACC TCTTCGTGCCAGAATGCGGGTCTTTTGGGCGATAATGGTATTTTGCGCAAGGCGGGTTTCAAGGGATACGTCGTC AGCGATTGGGGTGCAACGCACGATTCTGCAGAAGTGAACGCGAACAATGGTTTGGACATGGAGCAACCCGGCGAT TACATCGTGGTCGGAGGAGGCGTCTTTGGTGGGCTCAAGGACGCAGTCAACAAGGGACGGGTTTCCCAGAGACGT CTCAACGAAATGGTCGCGCGAATTCTTGCTCCTTGGTACTACCTCGAACAAGATTCTCCTGACTATCCTGAGACC AATTTTGACGTTCAACGACGCGATGGATCTGGCCCCAAGAACCTTCGGGTCAACGTTCGCTCCGAAGAACACACC GCCCTCGCCAGGGAGATTGCCTCTGCTTCTTCAGTCCTGTTGAAGAACGCCCGCTCCACCACCGACGGTACCCCT TCCGGAATCACTACGCGCGGCTTGCCACTCTCTAAAGACAGAATCAAGTCGATCGCTGTCATTGGTCAAGATGCC CGTCAGCTGAACCAGAATTGTTCGGAAATGGGTGAATGTAACGATGGCACCATGGTCGTTGGCTGGGGTTCTGGC TCTATCAACTTTGATCATGTCGTCCCTCCCATCGACGCCATCAAGTCTTTCACCAGTGACACTGCTACCATCACT GAGTCTCTAACGAACAACTTGGATGACGGTGTGCGGGCTGCGCGAGGCAAGGATGCTGCTCTTGTCTTTGTGAAC GCAGCGAGCGGTGAACTTGCCTTCTACACCGTCGTCCACGGAAACATGGGTGACAGGAATGACCTTGACCTTTGG TGGAGGGGTGGTAGCTTGATCGAGCGTGTAGCTGCTGTCAACAACAACACTATCGTCGTCGTTCATTCTGTTGGA CCCGTCTACATGCCCTGGAGCACCCATCCTAATATCACTGGTATCATTTACGCTGGCGCCCCTGGAGAGCAGACT GGACCTTCGATCGTCGACATCCTCTACGGAAACTACAACCCCAGTGGCAAGCTTCCTTTCAGCATTGCCGATAAC GAAGCTGCATACGGAACGACCATCGTTTACAACAGCCTTGGCTTCCCGGACATCGACTTCACTGAGAAGCTCCTC CTCGACTACCGATACATGGATGAGAAGAACATCCGCCCACGCTTCGAATTCGGTTTCGGTCTCTCCTACACCACC TTTGGATACTCCAACCTCGCTATCACTCGTGATGGCCCAGGATTCCAGGTTTCGTTCACTGTGGCCAACACTGGC GCATTTGCTGGTGCTGAGAAGCCTCAACTCTATCTTGCTTACCCTCCCAATGCTGGCCAGCCTAAGAAGGTTTTG CGTGGATTCGAAGAGGCTATTTTGGAGGTTGGTGCTAGCACCCGTGTCACTATCAACCTTAGCGAGCGCGATATG AGCGTTTGGGATGTTCCCAGCCAAAGCTATGTCCGCCTCCCGGGTGTTTACACGATCTACGTGGGTGCCTCGTCG CTCGATACCCGTCTGCAAGGAACCGTCACTATCTAA |
| Length | 2211 |
Transcript
| Sequence id | CopciAB_373418.T0 |
|---|---|
| Sequence |
>CopciAB_373418.T0 GGGACACCATCATGCGTTCTTCCTTCTCATTACTTGTGTTGCTCGCCATTCTCCAGGGCACTGTAGCCAGGAGAA CATGGGACGAAGCTTATGACCTAGCCAACAATGTGGTGAGCCAGATGACCTTGGACGAGAAGTTGGGAATCGTCA GGGGCACAGGCCAACTCAACCCTAACCGTCGTTGTGTTGGTGATACCAAGGCTGTTCCCCGACTGGGCATCCCGT CAATCTGTTTCGCAGATGGCCCCGCAGGAATGCGTCTCGTAAAAGAGACGACCGGCTTCCCTACTGGCATTAACG CCGCAGCAACCTTCAGTCGTCGCCTCATGAGAGAGCGTGGTGAGGCTTTGGGTCGGGAGTTCCGAGGCAAAGGAT CACACGTATTCTTGGGCCCATCCGTGGACGTTATGAGGAACCCCAAGGGAGGCCGTGCTTGGGAAGCCTTTGGTC CTGATCCTTATCTGAACGGAGAAGGCGCTTATGAGACTATCATGGGCGTCCAGAGCGTTGGAGTTCAAGCCTGCA TCAAGCACCTCATCGGCAATACCCAGGAGCACTGGAGATATGGTTACAGCGCCAACATGGACGACAGAACACTTC ACGAGATGTACTTTTATCCATTCCTTCGAAGTATTGAGGCCGACGTCTCGTCCGTCATGTGCGCTTACAATCGCT TCAACGGCACCTCTTCGTGCCAGAATGCGGGTCTTTTGGGCGATAATGGTATTTTGCGCAAGGCGGGTTTCAAGG GATACGTCGTCAGCGATTGGGGTGCAACGCACGATTCTGCAGAAGTGAACGCGAACAATGGTTTGGACATGGAGC AACCCGGCGATTACATCGTGGTCGGAGGAGGCGTCTTTGGTGGGCTCAAGGACGCAGTCAACAAGGGACGGGTTT CCCAGAGACGTCTCAACGAAATGGTCGCGCGAATTCTTGCTCCTTGGTACTACCTCGAACAAGATTCTCCTGACT ATCCTGAGACCAATTTTGACGTTCAACGACGCGATGGATCTGGCCCCAAGAACCTTCGGGTCAACGTTCGCTCCG AAGAACACACCGCCCTCGCCAGGGAGATTGCCTCTGCTTCTTCAGTCCTGTTGAAGAACGCCCGCTCCACCACCG ACGGTACCCCTTCCGGAATCACTACGCGCGGCTTGCCACTCTCTAAAGACAGAATCAAGTCGATCGCTGTCATTG GTCAAGATGCCCGTCAGCTGAACCAGAATTGTTCGGAAATGGGTGAATGTAACGATGGCACCATGGTCGTTGGCT GGGGTTCTGGCTCTATCAACTTTGATCATGTCGTCCCTCCCATCGACGCCATCAAGTCTTTCACCAGTGACACTG CTACCATCACTGAGTCTCTAACGAACAACTTGGATGACGGTGTGCGGGCTGCGCGAGGCAAGGATGCTGCTCTTG TCTTTGTGAACGCAGCGAGCGGTGAACTTGCCTTCTACACCGTCGTCCACGGAAACATGGGTGACAGGAATGACC TTGACCTTTGGTGGAGGGGTGGTAGCTTGATCGAGCGTGTAGCTGCTGTCAACAACAACACTATCGTCGTCGTTC ATTCTGTTGGACCCGTCTACATGCCCTGGAGCACCCATCCTAATATCACTGGTATCATTTACGCTGGCGCCCCTG GAGAGCAGACTGGACCTTCGATCGTCGACATCCTCTACGGAAACTACAACCCCAGTGGCAAGCTTCCTTTCAGCA TTGCCGATAACGAAGCTGCATACGGAACGACCATCGTTTACAACAGCCTTGGCTTCCCGGACATCGACTTCACTG AGAAGCTCCTCCTCGACTACCGATACATGGATGAGAAGAACATCCGCCCACGCTTCGAATTCGGTTTCGGTCTCT CCTACACCACCTTTGGATACTCCAACCTCGCTATCACTCGTGATGGCCCAGGATTCCAGGTTTCGTTCACTGTGG CCAACACTGGCGCATTTGCTGGTGCTGAGAAGCCTCAACTCTATCTTGCTTACCCTCCCAATGCTGGCCAGCCTA AGAAGGTTTTGCGTGGATTCGAAGAGGCTATTTTGGAGGTTGGTGCTAGCACCCGTGTCACTATCAACCTTAGCG AGCGCGATATGAGCGTTTGGGATGTTCCCAGCCAAAGCTATGTCCGCCTCCCGGGTGTTTACACGATCTACGTGG GTGCCTCGTCGCTCGATACCCGTCTGCAAGGAACCGTCACTATCTAAAGAGTCTTGTGAGCTCTCGATACCTTAA CTAACCACTCTACCTATCTATCTTTCACTGTCTTCACTGATCTACATAATCCTAATCGAATTTGAAACTG |
| Length | 2320 |
Gene
| Sequence id | CopciAB_373418.T0 |
|---|---|
| Sequence |
>CopciAB_373418.T0 GGGACACCATCATGCGTTCTTCCTTCTCATTACTTGTGTTGCTCGCCATTCTCCAGGGCACTGTAGCCAGGAGTA CGCTTTTCTGAATCCACAGTACGTTGTCTTATGCTAACGCCGTCTGTAGGAACATGGGACGAAGCTTATGACCTA GCCAACAATGTGGTGAGCCAGATGACCTTGGACGAGAAGTTGGGAATCGTCAGGGGCACAGGCCAACTCAACCCT AACCGTACGCCGGTCTCTGCAATGTCCAGGATCTGCCACTCACGCTGTTAACAGGTCGTTGTGTTGGTGATACCA AGGCTGTTCCCCGACTGGGCATCCCGTCAATCTGTTTCGCAGATGGCCCCGCAGGAATGCGTCTCGTAAAAGAGA CGACCGGCTTCCCTACTGGCATTAACGCCGCAGCAACCTTCAGTCGTCGCCTCATGAGAGAGCGTGGTGAGGCTT TGGGTCGGGAGTTCCGAGGCAAAGGATCACAGTGAGTTCGCCTGCGCATTTGTGGATTCCCACTCGGCTCATACT TCCCGCCGCAGCGTATTCTTGGGCCCATCCGTGGACGTTGTAAGTGGCTGCCCTGTAATTTGACCACTGGTACCC TCAATGTCGTGCATCTAGATGAGGAACCCCAAGGGAGGCCGTGCTTGGGAAGCGTACGTTCATCCTTTCACTCTA AATATAGCCCTTCGTCTAAATCCTCGTGTGCAGCTTTGGTCCTGATCCTTATCTGAACGGAGAAGGCGCTTATGA GACTATCATGGGCGTCCAGAGCGTTGGAGTTCAAGCCTGCATCAAGCACCTCATCGGCAATACCCAGGAGCACTG GAGATATGGTTACAGCGCCAACATGGACGACAGAACACTTCACGAGATGTACTTTTATCCATTCCTTCGAAGTAT TGAGGCAAGCAGGGAACCTCGTCTGGTCAAGGCAACTGGTGTTAACTAACTGATTATTTGTAGGCCGACGTCTCG TCCGTCATGTGCGCTTACAATCGCTTCAACGGCACCTCTTCGTGCCAGAATGCGGGTCTTTTGGGCGATAATGGT ATTTTGCGCAAGGCGGGTTTCAAGGGATACGTCGTCAGCGATTGGGGTGCAACGCACGATTCTGCAGAAGTGAAC GCGAACAATGGTTTGGACATGGAGCAACCCGGCGATTACATCGTGGTCGGAGGAGGCGTCTTTGGTGGGCTCAAG GACGCAGTCAACAAGGGACGGGTTTCCCAGAGAGTAAGTGGCATGACCTCCTCTGACCGGCTTGTTACTAACCTA TTTTCAGCGTCTCAACGAAATGGTCGCGCGAATTCTTGCTCCTTGGTACTACCTCGAACAAGATTCTCCTGTAAG TAATGGATTGGCCAACCTTCTTCAATGGACTTGAAGCTCAATTTACGCCTGTAGGACTATCCTGAGACCAATTTT GACGTTCAACGACGCGATGGATCTGGCCCCAAGAACCTTCGGGTCAACGTTCGCTCCGAAGAACACACCGCCCTC GCCAGGGAGATTGCCTCTGCTTCTTCAGTCCTGTTGAAGAACGCCCGCTCCACCACCGACGGTACCCCTTCCGGA ATCACTACGCGCGGCTTGCCACTCTCTAAAGACAGAATCAAGTCGATCGCTGTCATTGGTCAAGATGCCCGTCAG CTGAACCAGAATTGTTCGGAAATGGGTGAATGTAACGATGGCACCATGGTCGTTGGGTATGTAGGAATCTCCATT CTGTCGAGCAATTCGTTGACACTCATACTTAGCTGGGGTTCTGGCTCTATCAACTTTGATCATGTCGTCCCTCCC ATCGACGCCATCAAGTCTTTCACCAGTGACACTGCTACCATCACTGAGTCTCTAACGAACAACTTGGATGACGGT GTGCGGGCTGCGCGAGGCAAGGATGCTGCTCTTGTCTTTGTGAACGCGTAAGTTGCACCTCACAGATGTAGTGAA ACCAGCTGACAGGGCCCCTCAGAGCGAGCGGTGAACTTGCCTTCTACACCGTCGTCCACGGAAACATGGGTGACA GGAATGACCTTGACCTTTGGTGGAGGGGTGGTAGCTTGGTTAGTTTAATTTTGCTGTTCGTCCCAAGCTAATACT AAACTCACACTGGTGTAGATCGAGCGTGTAGCTGCTGTCAACAACAACACTATCGTCGTCGTTCATTCTGTTGGA CCCGTCTACATGCCCTGGAGCACCCATCCTAATATCACTGGTATCATTTACGCTGGCGCCCCTGGAGAGCAGACT GGACCTTCGATCGTCGACATCCTCTACGGAAACTACAACCCCAGTGGCAAGCTTCCTTTCAGCATTGCCGATGTA CGTCGCTTCCCTCTCGCTGCTTTGACCTTTGACTAAAGATGGTTTCTTTCACAGAACGAAGCTGCATACGGAACG ACCATCGTTTACAACAGCCTTGGCTTCCCGGACATCGACTTCACTGAGAAGCTCCTCCTCGACTACCGATACATG GATGAGAAGAACATCCGCCCACGCTTCGAATTCGGTTTCGGTCTCTCCTACACCACCTTTGGATACTCCAACCTC GCTATCACTCGTGATGGCCCAGGATTCCAGGTTTCGTTCACTGTGGCCAACACTGGCGCATTTGCTGGTGCTGAG AAGCCTCAACTCTATCTTGCTTACCCTCCCAATGCTGGCCAGCCTAAGAAGGTTTTGCGTGGATTCGAAGAGGCT ATTTTGGAGGTTGGTGCTAGCACCCGTGTCACTATCAACCTTAGCGAGCGCGATATGAGGTGCGTTCTTCGTGTT GCTTTTAGAGATGATCCGGAAGTTGATGGCATCCTTTGTAGCGTTTGGGATGTTCCCAGCCAAAGCTATGTCCGC CTCCCGGGTGTTTACACGATCTACGTGGGTGCCTCGTCGCTCGATACCCGTCTGCAAGGAACCGTCACTATCTAA AGAGTCTTGTGAGCTCTCGATACCTTAACTAACCACTCTACCTATCTATCTTTCACTGTCTTCACTGATCTACAT AATCCTAATCGAATTTGAAACTG |
| Length | 3023 |